RNA
Ribonucleic acid, or RNA, is a long, single-stranded chain of nucleic acid cells that contains proteins. Phosphate groups and the sugar ribose make up the backbone of RNA molecules (Lambert, 2001). RNA is an example of a cell chain that transports the genetic information of numerous viruses from the cell to the cytoplasm. All living cells include the macromolecule RNA, which is critical to the functioning of the cell. Aside from coding, decoding, regulation, and expression of genes, DNA holds the genetic instructions necessary for life’s growth and maintenance, and the enzyme responsible for synthesizing proteins carries those messages (Lambert, 2001). For example, mRNA helps to transmit genetic information that guides the production of specific proteins in cells.
Difference between RNA and DNA.
DNA contains a long chain of nucleotides of type B helix, whereas RNA has a short chain of nucleotides of type A helix as its structural foundation (Wang, 1995). Deoxyribose sugar, a kind of sugar lacking an oxygen atom, is found in DNA, whereas ribose is found in RNA. When it comes to genetic information, RNA only transfers the genetic code from the nucleus to generate protein, while DNA preserves genetic information for a lengthy period of time. In alkaline environments, DNA is more stable and less reactive than RNA, which is more reactive and fragile.
Difference between RNA dependent RNA polymerase and DNA dependent RNA polymerase.
A versatile and crucial enzyme in RNA virus genome replication and transcription, RdRp polymerase (or RdRp replicase) is an RNA replication enzyme that employs RNA templates to catalyze RNA replication (Arias et al., 2006). DNA-dependent RNA polymerase (DdRP or RNAP) is a key transcription enzyme in the replication systems of prokaryotic and eukaryotic organisms, as well as cytoplasmic DNA viruses (Aravind et al., 2003). It synthesizes the RNA strand that corresponds to a given RNA template.
Types RNA Transcribed within a cell.
Messenger RNA (mRNA), is a form of single-stranded RNA that plays a crucial role in transcription and is involved in protein synthesis. In order to ensure that the correct amino acids are integrated into the polypeptide chain during translation, the tRNA is accountable (Soler et al., 2014). The ribosomal RNA (rRNA) in eukaryotic cells serves as a framework for protein synthesis.
Structural differences of RNA types.
mRNA is characterized by a linear structure and is responsible for carrying the genetic information that is copied from DNA, while tRNA has a three-dimensional structure in the shape of an L. During the process of translation, it transports one amino acid at a time to the ever-expanding chain of a polypeptide (Flohr, 2017). Additionally, it is particular to each amino acid whereas ribosomal RNA molecules are spherical in shape and serve as a structural backbone for ribosomes.
Noncoding RNAs
Noncoding RNAs are molecules that are not translated into protein, and they are also large segments of the transcriptome that do not have apparent protein-coding roles. The pathophysiology of disease, for example, has been shown to benefit from the presence of noncoding RNAs in various biological processes.
Short and Long ncRNAs. Although they are only found in eukaryotes, short noncoding RNAs (ncRNAs) perform important roles in a wide variety of animals. There are long non-coding RNAs (lnc) in which the nucleotide length is at least 200 nucleotides (Ulitsky, 2018). Many functions, including the suppression of viruses and transposons, are regulated by various types of short RNAs, which are also responsible for numerous key developmental processes. For example, the interactions between the miRNAs and the mncRNAs, as well as the interactions between the mncRNAs and the mncRNAs at the transcriptional, post-transcriptional, translational, and post-translational stages; and at the translational level.
Introns
RNA transcripts or the DNA that encodes them contain non-coding regions known as introns. These sections are spliced away prior to translation into a protein. Protein-coding exons are found in the DNA (and RNA) of organisms. Introns, on the other hand, are the DNA/RNA that fills the gaps between exons. However, they are not coding for proteins, but rather vital for gene expression. It is the nucleotide sequences that carry an organism’s genetic information that constitute the genetic code (Chorev, & Carmel, 2012). Introns, from this perspective, function as hot places for recombination in the generation of new combinations of exons in the eukaryotic gene. To put it another way, we have them in our DNA because they were employed as a shortcut to assembling new genes during evolution.
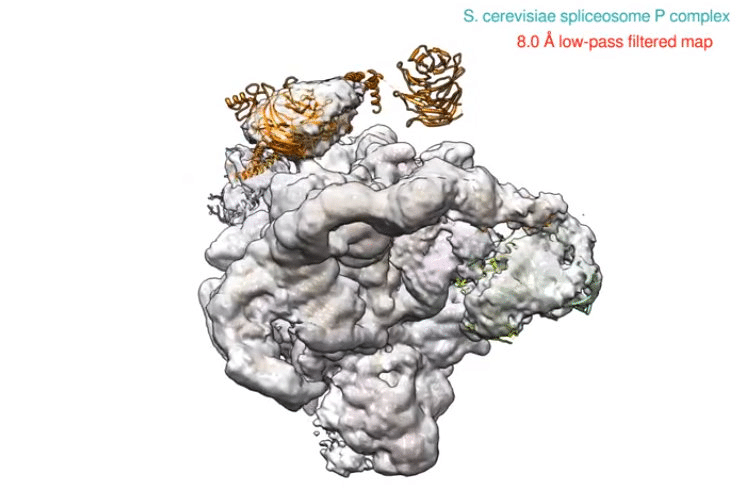
Spliceosomes
It is a huge RNA-protein complex that removes introns from the nucleus, a process known as splicing (Will, & Lührmann, 2011). As a result, messenger RNA is formed by removing the introns from precursor messenger RNA and joining the exons together in the cell nucleus.
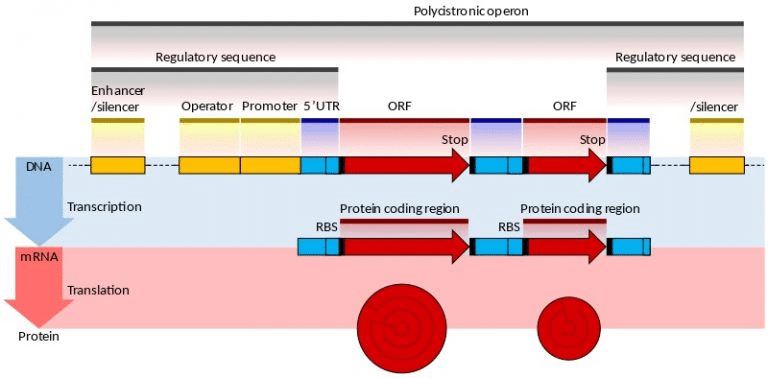
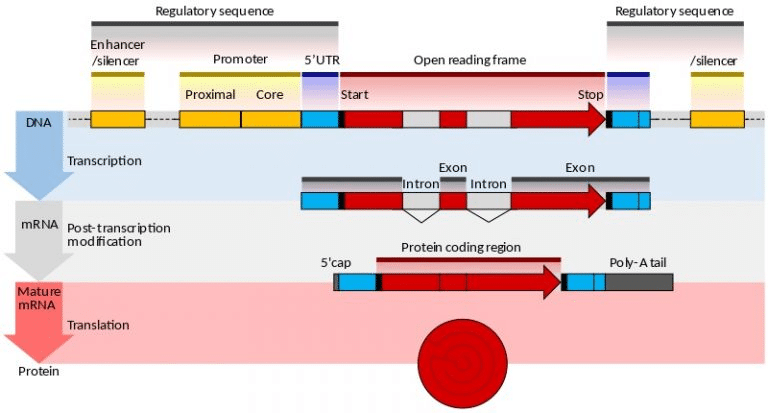
Deference between eukaryotic and procaryotic mRNA
The mRNA produced by prokaryotic operon transcription is known as prokaryotic mRNA, whereas the mRNA produced by eukaryotic single gene transcription is known as eukaryotic mRNA. When comparing mRNA from different organisms, it’s important to note that prokaryotic mRNA is polycistronic, while that from other organisms is monocistronic. An operon, in contrast to eukaryotic mRNA, has just a single gene and is composed of many structural genes. In prokaryotes, transcription is linked to translation, but transcription in eukaryotes occurs after transcription has finished in order to facilitate translation.
Role of Caping in Eukaryotic RNA.
In eukaryotic cells, the capping of mRNA 5′ ends is an essential structural modification because it facilitates translation, directs pre-mRNA splicing, and mRNA export from the nucleus, limits cellular 5′–3′ exonuclease degradation of mRNA, and allows recognition of foreign RNA (including viral transcripts) as ‘non-self’.
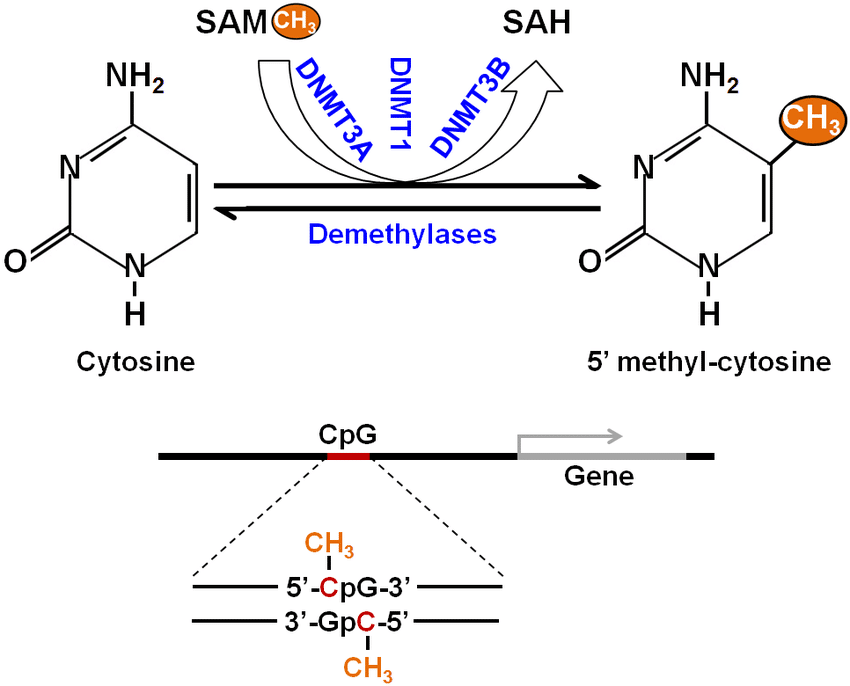
Methylation process during Caping
Protein lysine methyltransferases (PKMTs) and PRMTs are the most frequent enzymes that catalyze S-adenosylmethionine (SAM)-dependent methylation of protein substrates during caping. Cytosine can be modified either at its ring or at its exocyclic amino group to form 5-methylcytosine or N4 (Lui et al., 2014). N6-methyladenine is a product of adenine methylation.
RO
Role of 3’ and 5’ untranslated regions
3′-UTR influences the stability, exportability, and efficiency of translation of mRNA. The poly(A) tail, microRNA response elements (MREs), and AU-rich elements (AREs) are all present in this area. The 5′ UTR, which has been found to interact with proteins involved in metabolism, has been connected to biochemical activities. This region has also been shown to affect Drosophila’s sex-lethal gene. MRNA transport has been linked to the 5′ UTRs’ regulatory components as well.
Role of a poly tail in Eukaryotic mRNA.
In addition to protecting the mRNA molecule from enzymatic degradation, the poly (A) tail also aids in transcription termination, export of the mRNA from the nucleus, and translation of the mRNA. All except histone mRNAs in eukaryotic cells are polyadenylated, which is necessary for animal cell replication. The poly-A tail facilitates the export of the mature transcript from the nucleus to the ribosomes, where it is translated into a protein.
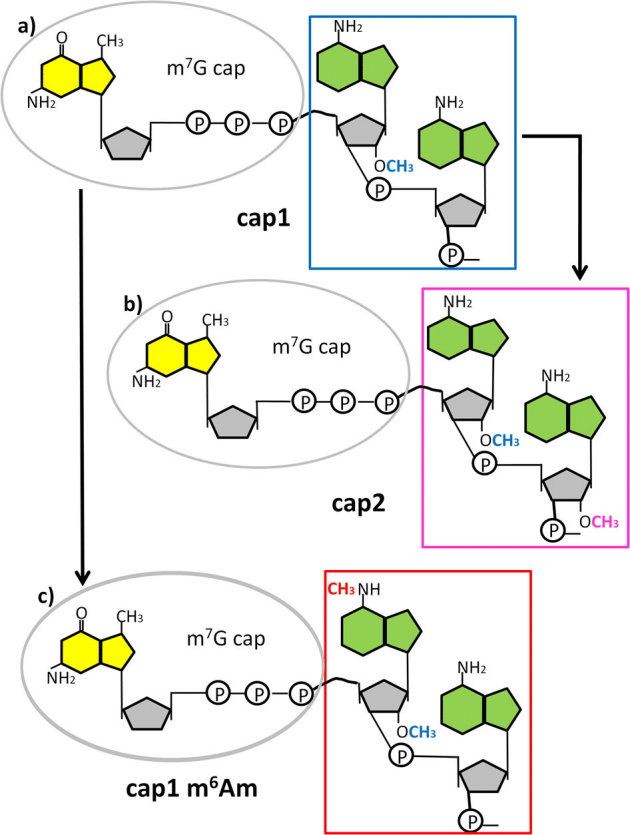
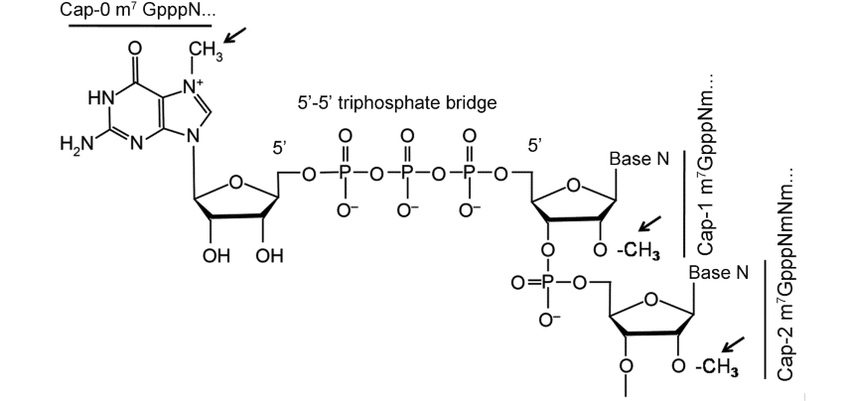
Deference between cap 0, cap 1, and cap 2 in Eukaryotic mRNA
Cap 0 consists of a N7-methyl guanosine that is triphosphate-linked to the 5′ nucleotide. The cap 0 structure is crucial for the effective translation of cap-carrying mRNA in the cell. Starting nucleotide p7GpppNm- is m7GpppNm-, where Nm indicates any nucleotide that has a 2′O methylation at the 2′O position. Cap 1 has been found to be essential for evasion of the innate immune response in cells. The shape of mRNA cap-1 could enhance expression during microinjection and transfection procedures. Cap 2 interacts with adenyl cyclase-associated protein and actin.
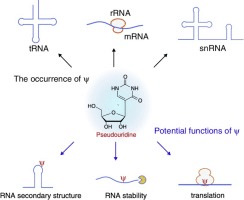
Role of Pseudo uridine in mRNA.
There are many benefits of Pseudouridine for mRNA stability and chemical properties. Pseudouridine-modified mRNA had a higher translatability than unmodified mRNA because unmodified mRNA is more likely to activate an RNA-dependent protein kinase (PKR) by binding. Translation efficiency decreases when the activity of this particular PKR leads to decreased levels of the translation initiation factor 2-alpha (eIF-2).
Advantages of using mRNA in making vaccines.
Because the mRNA vaccines are not constructed from an active virus, they are noninfectious (Thess 2012). They promote cellular and humoral immunity because antigens are generated inside cells (Thess et al., 2012). Because they do not include pathogen particles or inactivated pathogen, RNA vaccines are completely safe to use. As soon as the vaccination protein has been created, RNA strands in the vaccine are destroyed. Clinical trial data so far show that these vaccinations are effective and well-tolerated by healthy people, with just a small number of adverse reactions being reported. These vaccines can be developed quickly, giving them an edge over traditional vaccines that take longer to develop. Moderna’s mRNA-1273 vaccination for COVID-19 was developed in under two days. These products are also easier to standardize and can be produced with lower mistake rates, making them better suited to responding to large-scale outbreaks. The Pfizer–BioNTech vaccine was mass-produced in under 60 days instead of the customary 110 days required by traditional flu and polio vaccinations (before Pfizer began to streamline the manufacturing process). To put it into perspective, the entire process takes roughly 22 days: two weeks for DNA-based cloning and purification, four days for transcription from DNA to mRNA and four days for filling and finishing after the lipid nanoparticle-encapsulation step. In each production run, the majority of the days are devoted to meticulous quality control.
Advantages of using mRNA in gene therapy.
A number of features, including the ease of manipulation, the fast and transitory expression, and the adaptive conversion without mutagenesis, make MRNA-based therapy appropriate for dealing with the complexities and variations of glioma. Another advantage of mRNA over DNA is that its effects are transient rather than permanent. As a result, mRNA does not carry the risk of genomic integration, which is a real risk for DNA. mRNA has an inherent advantage over DNA-based therapies in terms of safety (Thess et al., 2012).
Messenger Vaccines RNA and how it works
The immunological response that is induced by a vaccine known as an mRNA vaccine is brought about by the administration of a replica of a molecule known as messenger RNA (mRNA).
How messenger RNA works. mRNA vaccines stimulate the production of certain proteins by acting as a transporter of protein information carried by mRNA. In the absence of vaccination, pathogens (such as viruses) or cancer cells would have to use the antigen-encoding mRNA that was provided to immune cells as a blueprint to manufacture a foreign protein. An adaptive immune response is activated by these protein molecules that directs the body to recognize and eliminate infections and cancer cells (Thess et al., 2012). Co-formulation of RNA with lipid nanoparticles protects and aids in the absorption of RNA strands into cells, resulting in mRNA delivery. The mRNA instructs cells to make the spike protein present on the surface of the SARS-CoV-2 virus in COVID-19 vaccinations (Kertes et al., 2022). To fight off what appears to be an infection caused by these foreign proteins, the body creates antibodies and other immune cells. The immune system is then prepared to launch an immediate reaction if it encounters the protein again in the future (such as in the case of a genuine SARS-CoV-2 viral infection).
Side Effects of DNA, RNA and protein-based Vaccines
Despite the fact that DNA/RNA vaccines were thoroughly tested for safety and efficacy before being widely distributed, a wide range of side effects, ranging from the mild to the severe, have been reported after immunization. Vaccination has been linked to viral infection, symptoms like headache and myalgia, as well as potentially fatal complications like myocardial damage and thrombosis (Kertes et al., 2022). There has to be more research on mRNA vaccines or the expressed protein antigens in order to develop safer vaccines, even though it is difficult to pinpoint the source of side effects in vaccine recipients.
Composition of Pfizer’s in messenger RNA and how it defers Moderna
Certainly, the mRNA in the Pfizer vaccine is not identical to that in the human body (Kertes et al., 2022). A cell is filled of defenses that are prepared to cut up RNA, especially any that does not belong there. Some of the mRNA building components have been replaced with “modified nucleosides” to prevent this from occurring.
The Moderna vaccine differs somewhat from the Pfizer vaccine in a few small ways. The first thing to notice is that the phrase. The mRNA-carrying fat bubbles have a slightly different chemical structure (Meo et al., 2021). In addition, the amount of RNA enclosed in the fat bubbles varies: In comparison to Pfizer, Moderna uses a substantially larger dosage (100 microgram) whereas Pfizer use 30 micrograms. As discussed above there is a lot about RNA and DNA that can be applied and be helping in life hence a lot of research is needed in this area.
References
Chorev, M., & Carmel, L. (2012). The function of introns. Frontiers in genetics, 3, 55.
Ferrer-Orta, C., Arias, A., Escarmís, C., & Verdaguer, N. (2006). A comparison of viral RNA-dependent RNA polymerases. Current opinion in structural biology, 16(1), 27-34.
Flohr, S. (2017). Identification and characterization of chemical modifications in mRNA of prokaryotes (Doctoral dissertation, The Weizmann Institute of Science (Israel)).
Gautheret, D., & Lambert, A. (2001). Direct RNA motif definition and identification from multiple sequence alignments using secondary structure profiles. Journal of molecular biology, 313(5), 1003-1011.
Iyer, L. M., Koonin, E. V., & Aravind, L. (2003). Evolutionary connection between the catalytic subunits of DNA-dependent RNA polymerases and eukaryotic RNA-dependent RNA polymerases and the origin of RNA polymerases. BMC structural biology, 3(1), 1-23.
Liu, J., & Jia, G. (2014). Methylation modifications in eukaryotic messenger RNA. Journal of genetics and genomics, 41(1), 21-33.
Liz, J., Portela, A., Soler, M., Gómez, A., Ling, H., Michlewski, G., … & Esteller, M. (2014). Regulation of pri-miRNA processing by a long noncoding RNA transcribed from an ultraconserved region. Molecular cell, 55(1), 138-147.
Meo, S. A., Bukhari, I. A., Akram, J., Meo, A. S., & Klonoff, D. C. (2021). COVID-19 vaccines: comparison of biological, pharmacological characteristics and adverse effects of Pfizer/BioNTech and Moderna Vaccines. Eur Rev Med Pharmacol Sci, 1663-1669.
Saciuk, Y., Kertes, J., Shamir Stein, N., & Ekka Zohar, A. (2022). Effectiveness of a third dose of BNT162b2 mRNA vaccine. The Journal of infectious diseases, 225(1), 30-33.
Schlake, T., Thess, A., Fotin-Mleczek, M., & Kallen, K. J. (2012). Developing mRNA-vaccine technologies. RNA biology, 9(11), 1319-1330.
Ulitsky, I. (2018). Interactions between short and long noncoding RNAs. FEBS letters, 592(17), 2874-2883.
Wang, S., & Kool, E. T. (1995). Origins of the large differences in stability of DNA and RNA helixes: C-5 methyl and 2′-hydroxyl effects. Biochemistry, 34(12), 4125-4132.
Will, C. L., & Lührmann, R. (2011). Spliceosome structure and function. Cold Spring Harbor perspectives in biology, 3(7), a003707.